In part 1 on this topic I showed this plot:
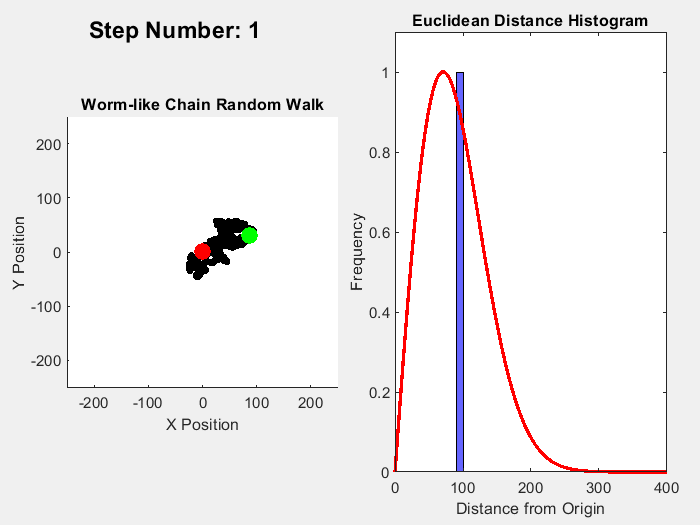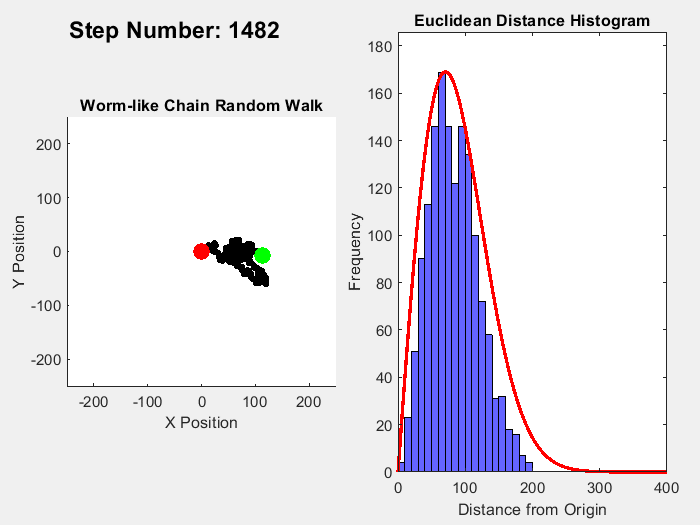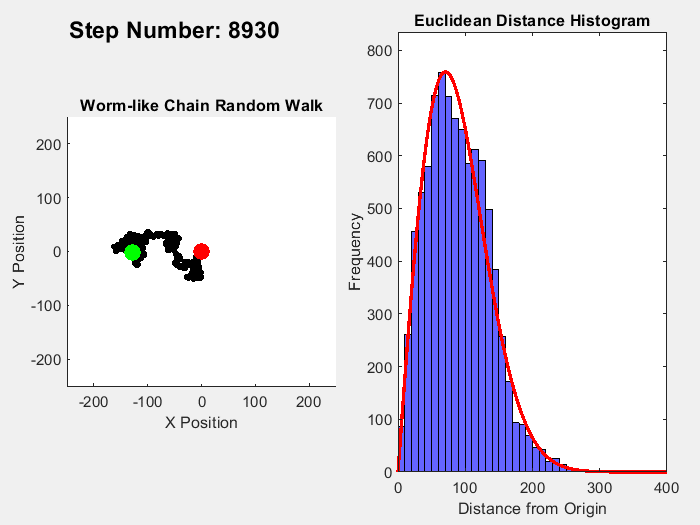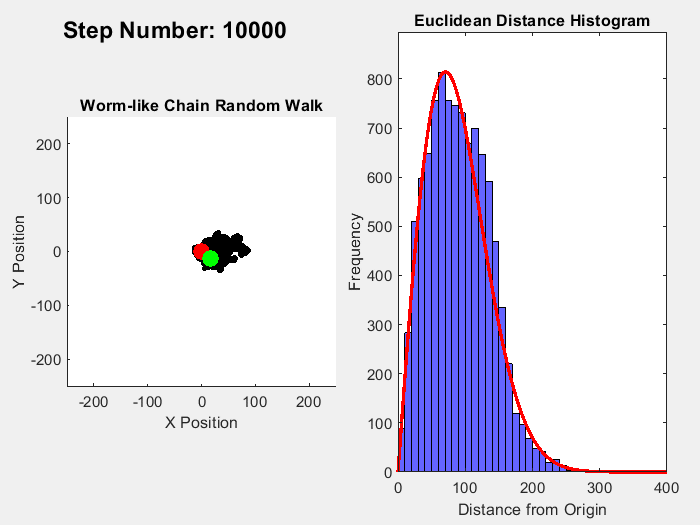
It's easy enough to reparametrize the radial coordinate in a way that flattens out the distribution. What is not obviously clear is what happens to the angular distributions of the individual chain links. In the plot shown, the distribution of the total chain length arises from a sum over 10,000 randomly oriented links. For each random link, the angle between the previous link and the following one is sampled from a uniform distribution.
On the other hand, reparametrizing the radial coordinate makes the total length uniformly distributed. As a consequence, the angular uniform distribution of the links must become distorted, non-uniform.
This fact, as we've also seen in the previous post, goes to the heart of what randomness is. When there are different parameters, there are multiple choices for which parameter to make "random": in our case, that can be either the total length distribution, or the radial distribution of the individual links. This is a great example of Bertrand's paradox.
So, let's derive the angular distributions of the chain links.
Wormlike-chain uniformized
The endpoint of link is located at
with in our case, and the radial distribution is
The full chain endpoint is , with distance from the origin, as plotted above, .
Coordinate system A1
In coordinate system A1, the link angles are sampled uniformly,
The resulting radial distribution is not exactly Rayleigh, because the endpoint cannot lie farther than . The exact finite- law is the 2D Pearson random-walk law:
So for the full chain we have
For large , this is extremely well approximated by the usual Rayleigh form
but the exact expression above is the right one if we want the support to end at , as we otherwise introduce avoidable artifacts.
Because the walk is isotropic, the polar angle of is uniform. So the position distribution of the endpoint of link is
The azimuth is flat, while the radius follows the Pearson law appropriate to steps.
Flattening the chain length distribution
To uniformize the endpoint radius of the full chain, the correct transformation is the cumulative distribution function
This maps the interval to , and by construction makes uniformly distributed.
Using the Bessel identity , the same CDF can be written as
This is the finite-support replacement for the large- approximation
To define a single coordinate change for the whole chain, not just for the final endpoint, we apply the same radial map to every point in the plane:
This gives us coordinate system A2.
Coordinate system A2
Under this map, the radial coordinate of every point becomes
while the polar angle stays the same. So the endpoint of link is still isotropic in angle, but its radial law changes by the Jacobian:
where .
So the full position distribution of the endpoint of link in A2 is
For , this becomes
which was the whole point of the construction.
So the transformation flattens the final endpoint distribution exactly, but it does not flatten the position distribution of every intermediate link endpoint. Each link index keeps its own nontrivial radial profile.
Angle laws in A1
There are two natural angles one can talk about.
The first is the absolute orientation of a link tangent in the lab frame. In A1 this is just , and by assumption
The second is the angle a link makes relative to the local radial direction. If the endpoint of link sits at polar angle , then the next link angle relative to the outward radial direction is
Since is sampled independently and uniformly, this is also uniform:
So in A1 the link tangent does not prefer radial or tangential directions. Locally, every direction is equally likely.
However, a radial coordinate change is not conformal unless it is just a linear rescaling. It stretches radial and tangential directions by different amounts.
Write a small displacement in polar form as
Under the map , this becomes
So radial and tangential directions scale by
Now suppose a link in A1 makes angle relative to the local radial direction. Its transformed angle in A2 satisfies
Define
Then
Since was uniform in A1, the transformed conditional angle law in A2 is
If , nothing happens and the angle law stays flat. But generically , so the local angle distribution is no longer uniform. Depending on radius, the transformation will favor directions closer to radial or closer to tangential.
The unconditional angle law for link is then obtained by averaging over the position distribution of the endpoint of link :
Equivalently, in the transformed radial coordinate,
So, taken together:
- in A1, link positions follow the Pearson random-walk law, and link angles are locally uniform;
- in A2, the final endpoint radius has been flattened, each intermediate link endpoint acquires a transformed radial profile, and the local link-angle law becomes position-dependent and non-uniform.
Running the same logic in the opposite direction
The previous section started from a chain whose link angles were sampled uniformly, and then changed coordinates so that the endpoint radius became uniform.
We can now reverse that logic.
Instead of starting from the angle law, let us start from the coordinate density that we ended up with in A2, and now simply declare that density to be the native one. That gives a second pair of coordinate systems, which we can call B1 and B2.
Coordinate system B1
In B1, the full-chain radial coordinate is taken to be uniform from the start:
For the intermediate link endpoints, we take exactly the radial profiles obtained above in A2. If is the endpoint of link , with polar coordinates , then
where
So the full-chain endpoint is flat in , while the intermediate link endpoints keep the nontrivial transformed profiles inherited from A2.
Now consider the angle of link relative to the local radial direction at . Call it . In A2 we found that the local angle law is
with
In B1 we now take that very same density as the native angle law:
So in B1 the length distribution is uniform, and the local angle law is non-uniform.
At the level of coordinate densities, B1 is therefore identical to A2:
and
So if one only inspects the density in the displayed coordinates, B1 and A2 look the same.
Flattening the angle laws
Now perform the opposite reparametrization: instead of flattening the radial coordinate, flatten the local angle law.
The clean way to do this is to undo the distortion factor . In A2 we had
So the inverse map is
with the branch chosen so that the full angle on is tracked continuously.
Equivalently, one can write this as the conditional cumulative transform
By construction, this makes the transformed angle variable uniform:
This is the angular analogue of the radial flattening that took us from A1 to A2.
Coordinate system B2
In B2, the local angle law has been flattened, so the links are described by uniform tangent angles once again:
Once the link angles have been put back into uniform form, the chain endpoint distribution reverts to the original Pearson random-walk law. So for the endpoint of link ,
and in particular for the full chain,
Likewise the local angle law is again flat:
So at the level of density, B2 is identical to A1:
and
In other words, we can run the whole construction in reverse:
- A1 starts with uniform link angles and produces a non-uniform endpoint radius;
- A2 flattens that radius and distorts the local angle law;
- B1 takes that flattened-radius/non-uniform-angle density as the starting point;
- B2 then flattens the angle law and recovers the original Pearson radial distribution.
So based on density alone,
Energy and entropy of the matching pairs
If one computes entropy and effective energy directly from the local coordinate density, then the matching pairs above necessarily agree.
Take any displayed coordinate with normalized density . Define the differential entropy
and the corresponding effective energy landscape
where is an arbitrary additive constant.
The corresponding mean effective energy is
So both the entropy and the average effective energy are determined entirely by the local density .
But we have already shown that the relevant coordinate densities match pairwise:
Therefore the corresponding entropies match:
and, with the same additive convention for , the mean effective energies match as well:
The same statement holds link-by-link for the endpoint distributions of every intermediate link , since those coordinate densities also match pairwise.
How to tell the two constructions apart
At this point, something subtle has happened.
We have constructed two different stories:
- in A1, the links are sampled with uniform angles, and the non-uniform radial distribution follows from that;
- in B1, the radial distribution is taken to be uniform from the start, and the non-uniform angle law follows from that.
After the corresponding coordinate changes, these can be made to look pairwise identical at the level of the displayed density:
So if all we are shown is the equilibrium density in the chosen coordinates, there is no obvious label attached saying which one came first.
This is exactly the same structural issue that appears in Bertrand’s paradox.
In Bertrand’s paradox, the phrase "choose a random chord" is incomplete until one specifies what is being sampled uniformly. Different choices of what counts as the primitive random variable lead to different quantitative answers, even though each one sounds perfectly reasonable in words.
The same thing is happening here. The phrase "make the chain random" is also incomplete until one decides what is being sampled uniformly.
One can start from
which says that the primitive randomness lies in the link angles, or one can start from
which says that the primitive randomness lies in the endpoint length.
These are quantitatively different ensembles. But by reparametrizing the coordinates, they can be made to share the same density.
So how do we tell them apart?
1: Correlations between variables
A single equilibrium density is too little information.
Suppose is some second observable: a bend angle, the angle of a link relative to the end-to-end direction, a local curvature, or anything else extracted from the chain.
In ensemble A, the joint law is
In ensemble B, the reweighting only depends on , so
This means the joint distributions are different whenever and are not independent.
So one straightforward way to distinguish the two constructions is to measure a joint density such as
rather than only the one-dimensional marginal of .
That one extra variable is enough to break the ambiguity.
2: Dynamical measurements
A still stronger way to distinguish the two is to stop looking only at equilibrium snapshots and instead record trajectories.
The reason is simple: a coordinate transformation changes how a given trajectory is described, but it does not create a new ensemble of trajectories. By contrast, changing the underlying sampling rule does.
In A, the path measure is generated by uniform angle sampling. In B, the path measure is reweighted by the final endpoint radius. So although one can make the equilibrium densities match in suitable coordinates, the trajectory statistics need not match.
That means one can compare quantities such as
or the response to an external perturbation. These depend on the full trajectory ensemble, not just the static equilibrium density, and so they generally distinguish the two constructions.
3: Measurements under different conditions
Another way to separate the two is to change a control parameter and ask whether the same underlying model continues to fit.
A passive coordinate transformation simply rewrites the same ensemble. It does not invent a new physical dependence on temperature or on the external controls.
By contrast, if one insists on imposing a given target density as fundamental, the effective weighting needed to maintain that target can itself acquire nontrivial dependence on the control parameter.
So if the same microscopic model is required to explain data across a family of conditions, the ambiguity can be broken.
What this has to do with Bertrand's paradox
Bertrand’s paradox is not just a trick about circles. It is a warning that the phrase "uniformly random" has no meaning until one specifies the measure.
That is exactly the moral here.
There is no contradiction between saying
- "the links are uniformly random, therefore the endpoint radius is non-uniform,"
and saying
- "the endpoint radius is uniformly random, therefore the link angles are non-uniform."
Both are mathematically legitimate once the primitive random variable has been specified.
What changes is the ensemble.
In that sense, A and B are different answers to the question: which variable is taken to be uniformly random before anything else is derived?
That is why the two constructions are quantitatively different, just as the different chord-generating procedures in Bertrand’s paradox are quantitatively different.
At the same time, within each construction, the paired coordinate systems are valid reparametrizations of one another:
So there are really two separate layers:
- the choice of ensemble, which determines what is taken as primitive randomness;
- the choice of coordinates, which determines how that ensemble is displayed.
Bertrand’s paradox lives in the first layer. The flattening transformations live in the second.
Once those two layers are separated, the structure becomes clearer:
- changing coordinates can make two different ensembles look the same at the level of a one-dimensional density;
- but looking at joint observables, unconditional angle laws, trajectories, or parameter dependence reveals that the ensembles are not actually the same.
Markovianity as a guide to interpretation
There is one more principle that is useful here, because it helps distinguish a good description from one that is merely a clever reparametrization.
That principle is Markovianity.
A variable is Markovian if its future depends only on its present value, not on the rest of its history:
In practice, this means that once the current state is known, the past adds no further predictive power.
Why does this matter here?
Because equilibrium densities alone are too permissive. A great many different microscopic constructions can be made to reproduce the same static distribution in some chosen coordinate. But the dynamics are much less forgiving. If the coordinate being used is a good one, then its evolution should be close to Markovian. If it is a bad one, then hidden variables will keep leaking through, and the apparent dynamics will remember the past.
This makes Markovianity a useful guide for interpretation.
If one coordinate system gives a description in which the observed dynamics are approximately Markovian, while another requires long memory kernels, path history, or extra hidden coordinates to explain the same data, then the former is usually the more natural description.
In that sense, Markovianity does not tell us which variable is "truly random" in some metaphysical sense. But it does tell us which parametrization is closer to being dynamically complete.
This is especially relevant in the present setting. A coordinate transformation can flatten a density, or flatten an angle law, or make one ensemble look deceptively similar to another. But if that transformation pushes important information into hidden correlations, then the resulting coordinate will generally look less Markovian.
So although equilibrium density alone cannot distinguish the constructions above, the time evolution often can. A good coarse-grained coordinate is not just one with a neat-looking stationary distribution. It is one in which the dynamics close on themselves as much as possible.